9 0 6 7 6
论文已发表
注册即可获取德孚的最新动态
IF 收录期刊
- 2.6 Breast Cancer (Dove Med Press)
- 3.9 Clin Epidemiol
- 3.3 Cancer Manag Res
- 3.9 Infect Drug Resist
- 3.6 Clin Interv Aging
- 4.8 Drug Des Dev Ther
- 2.8 Int J Chronic Obstr
- 8.0 Int J Nanomed
- 2.3 Int J Women's Health
- 3.2 Neuropsych Dis Treat
- 4.0 OncoTargets Ther
- 2.2 Patient Prefer Adher
- 2.8 Ther Clin Risk Manag
- 2.7 J Pain Res
- 3.3 Diabet Metab Synd Ob
- 4.3 Psychol Res Behav Ma
- 3.4 Nat Sci Sleep
- 1.9 Pharmgenomics Pers Med
- 3.5 Risk Manag Healthc Policy
- 4.5 J Inflamm Res
- 2.3 Int J Gen Med
- 4.1 J Hepatocell Carcinoma
- 3.2 J Asthma Allergy
- 2.3 Clin Cosmet Investig Dermatol
- 3.3 J Multidiscip Healthc

通过生物信息学分析和 qRT-PCR 验证鉴定食管腺癌中的转录因子-microRNA 网络
Authors Chen D, Lu T, Tan J, Zhao K, Li Y, Zhao W, Li H, Wang Q, Wang Y, Wei L
Received 11 January 2019
Accepted for publication 25 February 2019
Published 18 April 2019 Volume 2019:11 Pages 3315—3326
DOI https://doi.org/10.2147/CMAR.S201274
Checked for plagiarism Yes
Review by Single-blind
Peer reviewers approved by Dr Cristina Weinberg
Peer reviewer comments 2
Editor who approved publication: Dr Chien-Feng Li
Purpose: The rapidly rising incidence of
esophageal adenocarcinoma (EAC), which is usually diagnosed late with a poor
prognosis, has become a growing problem. This study investigated the potential
transcription factor (TF)-related molecular mechanisms of EAC by using
bioinformatics analysis and qRT-PCR validation.
Methods: Expression
profile datasets for mRNAs (GSE92396, GSE13898, GSE26886 and GSE1420) and
miRNAs (GSE16456) were downloaded from the GEO database. Overlapping
differentially expressed genes (DEGs) and differentially expressed miRNAs
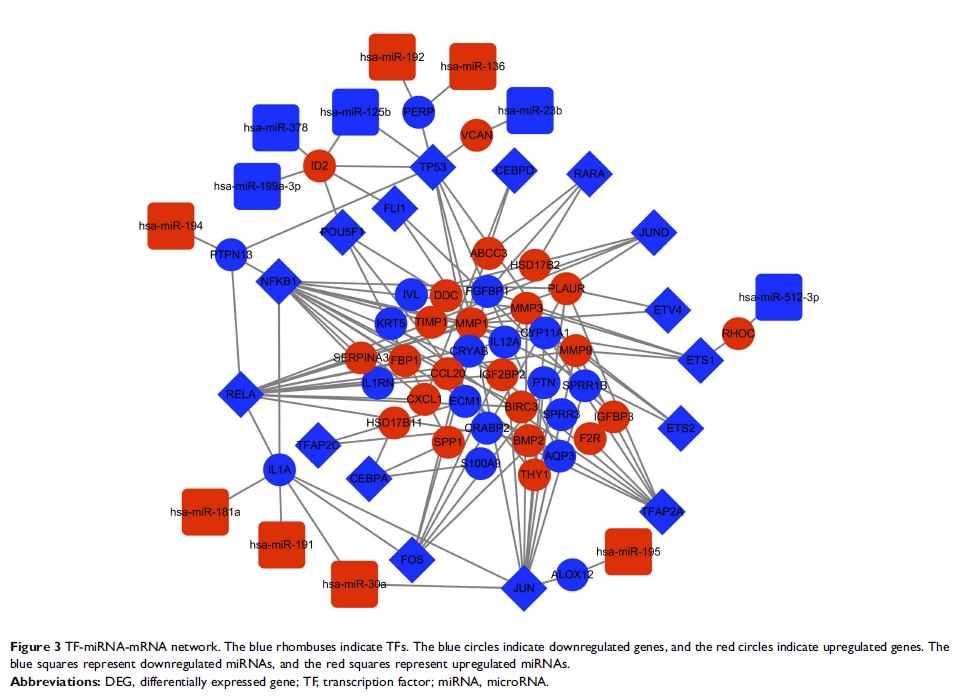
(DEMs) were identified through integrative analysis. Then, a TF-miRNA-mRNA
network was constructed based on bioinformatics data from the TRRUST, TRED and
miRTarBase database. Furthermore, overall survival analysis for the mRNAs and
miRNAs in the TF-miRNA-mRNA network was performed with data from TCGA, and
qRT-PCR was used to validate the results.
Results: A
total of 294 overlapping DEGs were identified in EAC tissues compared to normal
tissues, including 181 downregulated and 113 upregulated genes. Then, 16 TFs
that could target the DEGs and were related to cancer were predicted based on
public databases, and 41 DEGs that could be targeted were identified as key
genes. Additionally, 12 DEMs were predicted through miRTarBase to be associated
with the key genes, and TP53-(miR-125b)-ID2 and JUN-(miR-30a)-IL1A from the
TF-miRNA-mRNA network were identified to potentially play significant roles in
EAC. Furthermore, CCL20, IL1A, ABCC3, hsa-miR-23b, and hsa-miR-191, which are
involved in the TF-miRNA-mRNA network, were found to be significantly
associated with patient survival in EAC. Finally, the expression of a
miRNA-mRNA pair (hsa-miR-30a-5p and IL1A) was revealed to be correlated with
prognosis.
Conclusion: In
this study, a TF-miRNA-mRNA network was constructed to analyze the potential
molecular mechanisms of EAC. Key genes and miRNAs associated with patient
survival were identified, which may reveal promising approaches for EAC
diagnosis and therapy.
Keywords: esophageal
adenocarcinoma, prognosis, differentially expressed genes, microRNA,
transcription factor